
Made up of one old, conserved strand of DNA and one new one. RNA primers within the lagging strand are removed by the exonuclease activity of DNA polymerase I, and the Okazaki fragments are joined by DNA ligase. Replication is described as semi- conservative because each DNA molecule is On the leading strand, DNA is synthesized continuously, whereas on the lagging strand, DNA is synthesized in short stretches called Okazaki fragments. The Okazaki fragments are formed in the lagging strands, and the replication of the lagging strand is discontinuous. Finally the enzyme DNA ligase seals up theįragments of DNA in both strands to form a continuous double strand. The DNA ligase is the enzyme that forms a phosphodiester bond between the hydroxyl group of the carbon at the 3 end of one nucleotide in the Okazaki fragment with the phosphate group at the 5 end of the next Okazaki fragment. Another DNA polymerase enzyme then fills in the gaps The new DNA has been made the enzyme exonuclease removes all the RNA primersįrom both strands of DNA. The next primer is then added further down the lagging strand.Īnother Okazaki fragment is then made and the process is repeated again. DNA polymerase then adds a short row of DNA bases in the 5' In a series of small chunks called Okazaki fragments. The opposite direction the DNA polymerase can therefore only make this strand The lagging strand, cannot be made in this continuous way because it runs in Polymerase adding bases one by one in the 5' to 3' direction. New strands of DNA, the leading strand, is made continuously, the DNA Only add DNA bases in one direction, from the 5' end to the 3' end. Recent studies have implicated LIG3 complexed with XRCC1 as an. An enzyme called DNA polymeraseīinds to the primer and will make the new strand of DNA. DNA ligase 1 (LIG1) is known as the major DNA ligase responsible for Okazaki fragment joining. The construction of the new strand of DNA. 
Makes a small piece of RNA called a primer. This is the first direct evidence that the interaction between DNA ligase I and PCNA coordinates the synthesis and ligation of Okazaki fragments. An enzyme called primase starts the process. Recent findings reveal that Okazaki fragment maturation is highly coordinated. DNA ligase III : complexes with DNA repair protein XRCC1 to aid in sealing DNA during the process of nucleotide excision repair and recombinant fragments. The separated strands each provide a template for creatingĪ new strand of DNA. DNA ligase I: ligates the nascent DNA of the lagging strand after the Ribonuclease H has removed the RNA primer from the Okazaki fragments. Unzipping is done by an enzyme called helicase and results in the formation ofĪ replication fork. The first step in DNA replication is to separate the two strands. 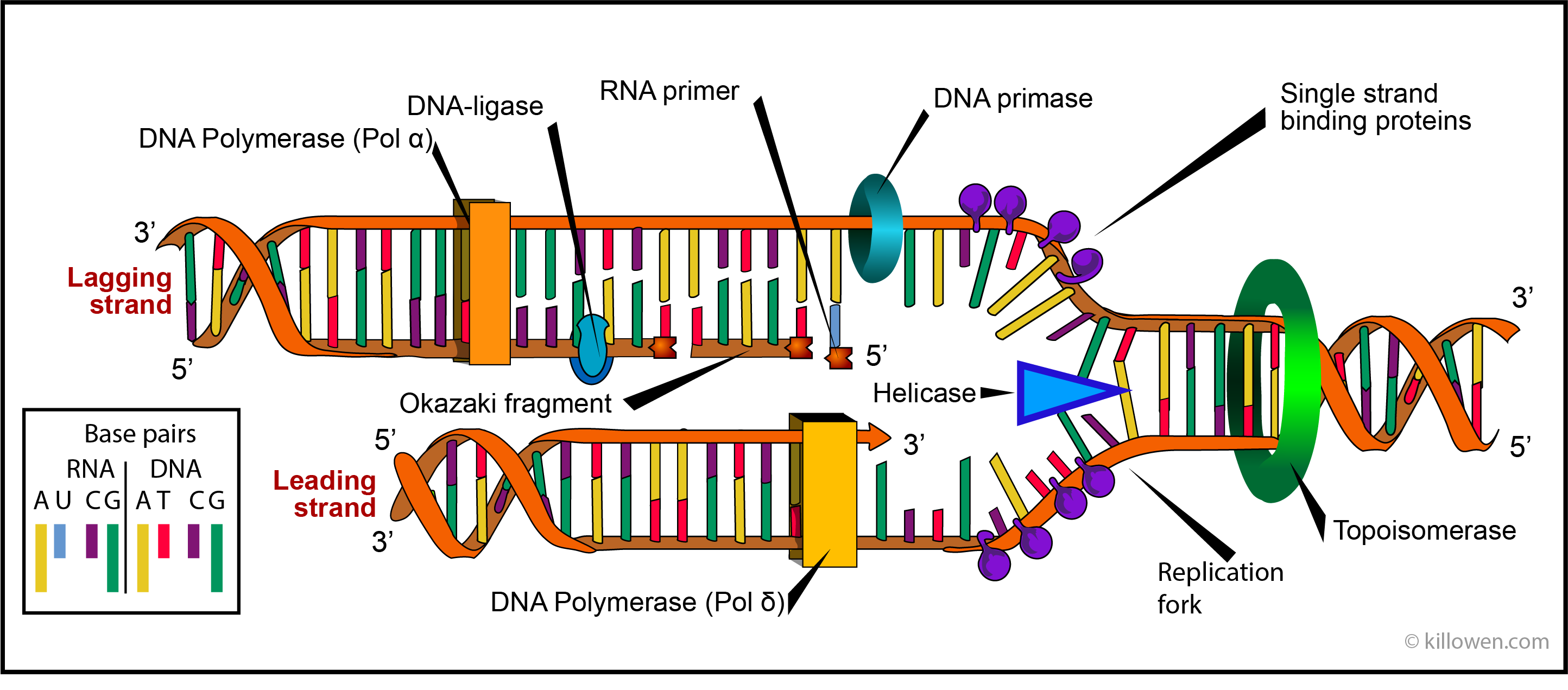
This determines how each strand of DNA is Will be in an A in the opposite strand, and wherever there's a C there will beĪ G in the other strand. This means that wherever there's a T in one strand there Each strand is made up a sequence ofįour chemical bases represented by the letters A, C, G and T. It shows how both strands of the DNA helix are unzipped and copied to produce two identical DNA molecules.ĭNA is a molecule made up of two strands twistedĪround each other in a double helix shape. This 3D animation shows you how DNA is copied in a cell.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
